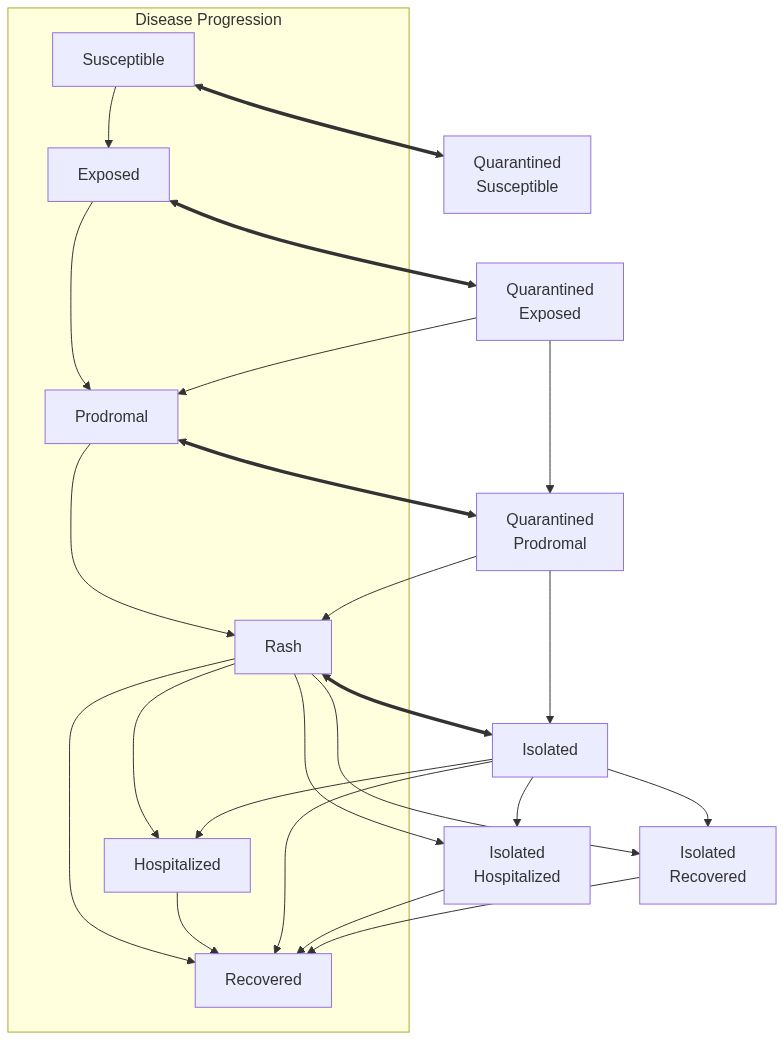
ModelMeaslesMixing creates a measles epidemiological model with mixing
between different population groups. The model includes vaccination,
quarantine, isolation, and contact tracing mechanisms.
Usage
ModelMeaslesMixing(
n,
prevalence,
contact_matrix,
vax_reduction_recovery_rate = 0.5,
transmission_rate = 0.9,
prop_vaccinated,
vax_efficacy = 0.97,
quarantine_period = 21,
quarantine_willingness = 1,
isolation_willingness = 1,
isolation_period = 4,
incubation_period = 12,
prodromal_period = 4,
rash_period = 3,
hospitalization_rate = 0.2,
hospitalization_period = 7,
days_undetected = 2,
contact_tracing_success_rate = 1,
contact_tracing_days_window = 4
)Arguments
- n
Number of individuals in the population.
- prevalence
Double. Initial proportion of individuals with the virus.
- contact_matrix
A numeric square matrix with the expected number of contacts per time step between population groups.
- vax_reduction_recovery_rate
Double. Vaccine reduction in recovery rate (default: 0.5).
- transmission_rate
Numeric scalar between 0 and 1. Probability of transmission (default: 0.9).
- prop_vaccinated
Double. Proportion of population that is vaccinated.
- vax_efficacy
Double. Vaccine efficacy rate (default: 0.99).
- quarantine_period
Integer. Number of days for quarantine (default: 21).
- quarantine_willingness
Double. Proportion of agents willing to quarantine (default: 1).
- isolation_willingness
Double. Proportion of agents willing to isolate (default: 1).
- isolation_period
Integer. Number of days for isolation (default: 4).
- incubation_period
Double. Duration of incubation period (default: 12).
- prodromal_period
Double. Duration of prodromal period (default: 4).
- rash_period
Double. Duration of rash period (default: 3).
- hospitalization_rate
Double. Rate of hospitalization (default: 0.2).
- hospitalization_period
Double. Period of hospitalization (default: 7).
- days_undetected
Double. Number of days an infection goes undetected (default: 2).
- contact_tracing_success_rate
Double. Probability of successful contact tracing (default: 1.0).
- contact_tracing_days_window
Integer. Number of days before rash onset that will be considered for contact tracing (default: 4).
Value
The
ModelMeaslesMixingfunction returns a model of classes epiworldR::epiworld_model and epiworld_measlesmixing.
Details
The contact_matrix is a square matrix of contact rates between entities.
Entry [i, j] gives the expected number of contacts that an agent in entity
i has with agents in entity j during a time step. The matrix should have
one row and one column per entity in the model.
The model includes three distinct phases of measles infection: incubation, prodromal, and rash periods. Vaccination provides protection against infection and may reduce recovery time.
The epiworldR::initial_states function allows the user to set the initial state of the model. In particular, the user can specify how many of the non-infected agents have been removed at the beginning of the simulation.
Hospitalization Probability
Instead of hospitalization probability, the model uses hospitalization rate. The following equation describes the hospitalization probability as a function of the hospitalization rate and recovery rate (from rash):
$$ P(\text{hospitalization}) = \frac{ \text{hospitalization}_\text{rate}}{ \text{hospitalization}_\text{rate} + \text{recovery}_\text{rate} } $$
Where the \(\text{recovery}_\text{rate}\) is given by the rash period (1/duration of it). In other words, to match a desired hospitalization probability, the user needs to use the following:
h_rate <- p_hosp * (1/rash_days) / (1 - p_hosp)See also
epiworld-methods
Other Models:
ModelMeaslesMixingRiskQuarantine(),
ModelMeaslesSchool()
Other measles models:
ModelMeaslesMixingRiskQuarantine(),
ModelMeaslesSchool()
Examples
# Start off creating three entities.
# Individuals will be distributed randomly between the three.
e1 <- entity("Population 1", 3e3, as_proportion = FALSE)
e2 <- entity("Population 2", 3e3, as_proportion = FALSE)
e3 <- entity("Population 3", 3e3, as_proportion = FALSE)
# Contact matrix including within- and between-group contact rates
cmatrix <- (c(
c(0.9, 0.05, 0.05),
c(0.1, 0.8, 0.1),
c(0.1, 0.2, 0.7)
) * 15) |> matrix(byrow = TRUE, nrow = 3)
N <- 9e3
measles_model <- ModelMeaslesMixing(
n = N,
prevalence = 1 / N,
transmission_rate = 0.9,
vax_efficacy = 0.97,
vax_reduction_recovery_rate = 0.8,
incubation_period = 10,
prodromal_period = 3,
rash_period = 7,
contact_matrix = cmatrix,
hospitalization_rate = 0.1,
hospitalization_period = 10,
days_undetected = 2,
quarantine_period = 14,
quarantine_willingness = 0.9,
isolation_willingness = 0.8,
isolation_period = 10,
prop_vaccinated = 0.95,
contact_tracing_success_rate = 0.8,
contact_tracing_days_window = 4
)
# Adding the entities to the model
measles_model |>
add_entity(e1) |>
add_entity(e2) |>
add_entity(e3)
set.seed(331)
run(measles_model, ndays = 100)
#> _________________________________________________________________________
#> Running the model...
#> ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| done.
summary(measles_model)
#> ________________________________________________________________________________
#> ________________________________________________________________________________
#> SIMULATION STUDY
#>
#> Name of the model : Measles with Mixing and Quarantine
#> Population size : 9000
#> Agents' data : (none)
#> Number of entities : 3
#> Days (duration) : 100 (of 100)
#> Number of viruses : 1
#> Last run elapsed t : 14.00ms
#> Last run speed : 62.34 million agents x day / second
#> Rewiring : off
#> Last seed used : 8019
#>
#> Global events:
#> - Quarantine process (runs daily)
#>
#> Virus(es):
#> - Measles
#>
#> Tool(s):
#> - MMR
#>
#> Model parameters:
#> - (IGNORED) Vax improved recovery : 0.8000
#> - Contact tracing days window : 4.0000
#> - Contact tracing success rate : 0.8000
#> - Days undetected : 2.0000
#> - Hospitalization period : 10.0000
#> - Hospitalization rate : 0.1000
#> - Incubation period : 10.0000
#> - Isolation period : 10.0000
#> - Isolation willingness : 0.8000
#> - Prodromal period : 3.0000
#> - Quarantine period : 14.0000
#> - Quarantine willingness : 0.9000
#> - Rash period : 7.0000
#> - Transmission rate : 0.9000
#> - Vaccination rate : 0.9500
#> - Vax efficacy : 0.9700
#>
#> Distribution of the population at time 100:
#> - ( 0) Susceptible : 8999 -> 8998
#> - ( 1) Latent : 1 -> 0
#> - ( 2) Prodromal : 0 -> 0
#> - ( 3) Rash : 0 -> 0
#> - ( 4) Isolated : 0 -> 0
#> - ( 5) Isolated Recovered : 0 -> 1
#> - ( 6) Quarantined Latent : 0 -> 0
#> - ( 7) Quarantined Susceptible : 0 -> 0
#> - ( 8) Quarantined Prodromal : 0 -> 0
#> - ( 9) Quarantined Recovered : 0 -> 0
#> - (10) Hospitalized : 0 -> 0
#> - (11) Recovered : 0 -> 1
#>
#> Transition Probabilities:
#> - Susceptible 1.00 0.00 - - - - - - - - - -
#> - Latent - 0.60 0.20 - - - 0.20 - - - - -
#> - Prodromal - - - 1.00 - - - - - - - -
#> - Rash - - - - - 1.00 - - - - - -
#> - Isolated - - - - 0.83 - - - - - 0.17 -
#> - Isolated Recovered - - - - - 1.00 - - - - - -
#> - Quarantined Latent - - - - - - 0.67 - 0.33 - - -
#> - Quarantined Susceptible - - - - - - - - - - - -
#> - Quarantined Prodromal - - - - 1.00 - - - - - - -
#> - Quarantined Recovered - - - - - - - - - - - -
#> - Hospitalized - - - - - - - - - - 0.96 0.04
#> - Recovered - - - - - - - - - - - 1.00
#>