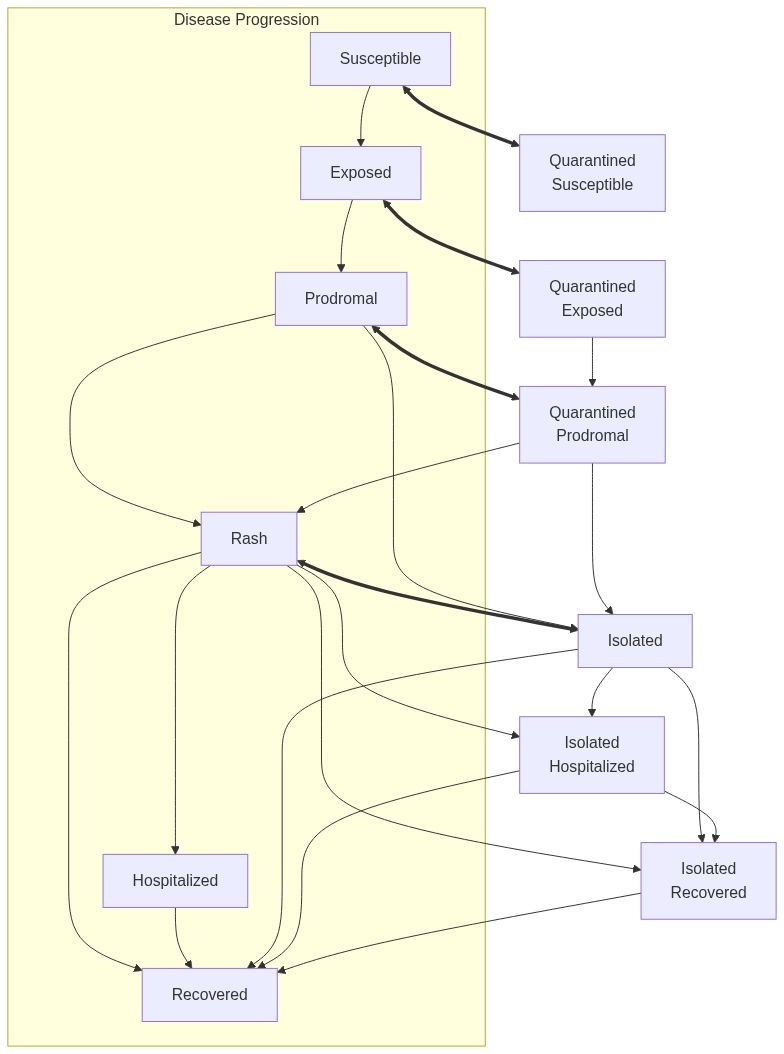
Measles model with mixing and risk-based quarantine
Source:R/ModelMeaslesMixingRiskQuarantine.R
ModelMeaslesMixingRiskQuarantine.RdModelMeaslesMixingRiskQuarantine creates a measles epidemiological model with mixing
between different population groups and risk-based quarantine strategies. The model
includes vaccination, quarantine with three risk levels (high, medium, low), isolation,
and contact tracing mechanisms.
Usage
ModelMeaslesMixingRiskQuarantine(
n,
prevalence,
contact_matrix,
transmission_rate = 0.9,
prop_vaccinated,
vax_efficacy = 0.97,
quarantine_period_high = 21,
quarantine_period_medium = 14,
quarantine_period_low = 7,
quarantine_willingness = 1,
isolation_willingness = 1,
isolation_period = 4,
incubation_period = 12,
prodromal_period = 4,
rash_period = 3,
hospitalization_rate = 0.2,
hospitalization_period = 7,
days_undetected = 2,
detection_rate_quarantine = 0.5,
contact_tracing_success_rate = 1,
contact_tracing_days_window = 4
)Arguments
- n
Number of individuals in the population.
- prevalence
Double. Initial proportion of individuals with the virus.
- contact_matrix
A numeric square matrix with the expected number of contacts per time step between population groups.
- transmission_rate
Numeric scalar between 0 and 1. Probability of transmission (default: 0.9).
- prop_vaccinated
Double. Proportion of population that is vaccinated.
- vax_efficacy
Double. Vaccine efficacy rate (default: 0.99).
- quarantine_period_high
Integer. Number of days for quarantine for high-risk contacts (default: 21).
- quarantine_period_medium
Integer. Number of days for quarantine for medium-risk contacts (default: 14).
- quarantine_period_low
Integer. Number of days for quarantine for low-risk contacts (default: 7).
- quarantine_willingness
Double. Proportion of agents willing to quarantine (default: 1).
- isolation_willingness
Double. Proportion of agents willing to isolate (default: 1).
- isolation_period
Integer. Number of days for isolation (default: 4).
- incubation_period
Double. Duration of incubation period (default: 12).
- prodromal_period
Double. Duration of prodromal period (default: 4).
- rash_period
Double. Duration of rash period (default: 3).
- hospitalization_rate
Double. Rate of hospitalization (default: 0.2).
- hospitalization_period
Double. Period of hospitalization (default: 7).
- days_undetected
Double. Number of days rash goes undetected (default: 2).
- detection_rate_quarantine
Double. Detection rate of prodromal agents during active quarantine periods (default: 0.5).
- contact_tracing_success_rate
Double. Probability of successful contact tracing (default: 1.0).
- contact_tracing_days_window
Integer. Number of days before and after the onset of symptoms for which contact tracing is effective (default: 4).
Value
The
ModelMeaslesMixingRiskQuarantinefunction returns a model of classes epiworldR::epiworld_model and epiworld_measlesmixingriskquarantine.
Details
The contact_matrix is a square matrix of contact rates between entities.
Entry [i, j] gives the expected number of contacts that an agent in entity
i has with agents in entity j during a time step. The matrix should have
one row and one column per entity in the model.
The model includes three distinct phases of measles infection: latent (incubation), prodromal, and rash periods. Vaccination provides protection against transmission.
Risk-based quarantine strategies assign different quarantine durations based on exposure risk:
High Risk: Unvaccinated agents who share entity membership with the case
Medium Risk: Unvaccinated agents who contacted an infected individual but don't share entity membership
Low Risk: Other unvaccinated agents
The epiworldR::initial_states function allows the user to set the initial state of the model. In particular, the user can specify how many of the non-infected agents have been removed at the beginning of the simulation.
The model uses hospitalization rates instead of probabilities. To learn
more about this, see the documentation in ModelMeaslesMixing().
See also
epiworld-methods
Other Models:
ModelMeaslesMixing(),
ModelMeaslesSchool()
Other measles models:
ModelMeaslesMixing(),
ModelMeaslesSchool()
Examples
# Start off creating three entities.
# Individuals will be distributed randomly between the three.
e1 <- entity("Population 1", 3e3, as_proportion = FALSE)
e2 <- entity("Population 2", 3e3, as_proportion = FALSE)
e3 <- entity("Population 3", 3e3, as_proportion = FALSE)
# Contact matrix including within- and between-group contact rates
cmatrix <- c(
c(0.9, 0.05, 0.05),
c(0.1, 0.8, 0.1),
c(0.1, 0.2, 0.7)
) |> matrix(byrow = TRUE, nrow = 3) * 15
N <- 9e3
measles_model <- ModelMeaslesMixingRiskQuarantine(
n = N,
prevalence = 1 / N,
transmission_rate = 0.9,
vax_efficacy = 0.97,
incubation_period = 10,
prodromal_period = 3,
rash_period = 7,
contact_matrix = cmatrix,
hospitalization_rate = 0.1,
hospitalization_period = 10,
days_undetected = 2,
quarantine_period_high = 21,
quarantine_period_medium = 14,
quarantine_period_low = 7,
quarantine_willingness = 0.9,
isolation_willingness = 0.8,
isolation_period = 10,
prop_vaccinated = 0.95,
detection_rate_quarantine = 0.5,
contact_tracing_success_rate = 0.8,
contact_tracing_days_window = 4
)
# Adding the entities to the model
measles_model |>
add_entity(e1) |>
add_entity(e2) |>
add_entity(e3)
set.seed(331)
run(measles_model, ndays = 100)
#> _________________________________________________________________________
#> Running the model...
#> ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| done.
summary(measles_model)
#> ________________________________________________________________________________
#> ________________________________________________________________________________
#> SIMULATION STUDY
#>
#> Name of the model : Measles with Mixing and Risk-based Quarantine
#> Population size : 9000
#> Agents' data : (none)
#> Number of entities : 3
#> Days (duration) : 100 (of 100)
#> Number of viruses : 1
#> Last run elapsed t : 55.00ms
#> Last run speed : 16.29 million agents x day / second
#> Rewiring : off
#> Last seed used : 8019
#>
#> Global events:
#> - Update infected individuals (runs daily)
#>
#> Virus(es):
#> - Measles
#>
#> Tool(s):
#> - MMR
#>
#> Model parameters:
#> - Contact tracing days window : 4.0000
#> - Contact tracing success rate : 0.8000
#> - Days undetected : 2.0000
#> - Detection rate quarantine : 0.5000
#> - Hospitalization period : 10.0000
#> - Hospitalization rate : 0.1000
#> - Incubation period : 10.0000
#> - Isolation period : 10.0000
#> - Isolation willingness : 0.8000
#> - Prodromal period : 3.0000
#> - Quarantine period high : 21.0000
#> - Quarantine period low : 7.0000
#> - Quarantine period medium : 14.0000
#> - Quarantine willingness : 0.9000
#> - Rash period : 7.0000
#> - Transmission rate : 0.9000
#> - Vaccination rate : 0.9500
#> - Vax efficacy : 0.9700
#>
#> Distribution of the population at time 100:
#> - ( 0) Susceptible : 8999 -> 8466
#> - ( 1) Latent : 1 -> 14
#> - ( 2) Prodromal : 0 -> 1
#> - ( 3) Rash : 0 -> 3
#> - ( 4) Isolated : 0 -> 3
#> - ( 5) Isolated Recovered : 0 -> 5
#> - ( 6) Quarantined Latent : 0 -> 0
#> - ( 7) Quarantined Susceptible : 0 -> 388
#> - ( 8) Quarantined Prodromal : 0 -> 0
#> - ( 9) Quarantined Recovered : 0 -> 0
#> - (10) Hospitalized : 0 -> 9
#> - (11) Recovered : 0 -> 111
#>
#> Transition Probabilities:
#> - Susceptible 1.00 0.00 - - - - 0.00 0.00 - - - -
#> - Latent - 0.89 0.10 - - - 0.01 - 0.00 - - -
#> - Prodromal - - 0.68 0.15 0.17 - - - - - - -
#> - Rash - - - 0.39 0.37 0.10 - - - - 0.09 0.06
#> - Isolated - - - 0.04 0.73 0.11 - - - - 0.12 0.00
#> - Isolated Recovered - - - - - 0.85 - - - - - 0.15
#> - Quarantined Latent - 0.07 - - - - 0.76 - 0.17 - - -
#> - Quarantined Susceptible 0.03 - - - - - - 0.97 - - - -
#> - Quarantined Prodromal - - - - 0.60 - - - 0.40 - - -
#> - Quarantined Recovered - - - - - - - - - - - -
#> - Hospitalized - - - - - - - - - - 0.90 0.10
#> - Recovered - - - - - - - - - - - 1.00
#>