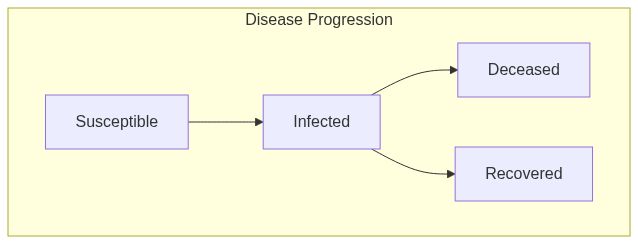
Susceptible Infected Removed Deceased model (SIRD connected)
Source:R/ModelSIRDCONN.R
ModelSIRDCONN.RdSusceptible Infected Removed Deceased model (SIRD connected)
Usage
ModelSIRDCONN(
name,
n,
prevalence,
contact_rate,
transmission_rate,
recovery_rate,
death_rate
)Arguments
- name
String. Name of the virus
- n
Number of individuals in the population.
- prevalence
Double. Initial proportion of individuals with the virus.
- contact_rate
Numeric scalar. Average number of contacts per step.
- transmission_rate
Numeric scalar between 0 and 1. Probability of transmission.
- recovery_rate
Numeric scalar between 0 and 1. Probability of recovery.
- death_rate
Numeric scalar between 0 and 1. Probability of death.
Value
The
ModelSIRDCONNfunction returns a model of class epiworld_model.
Details
The initial_states function allows the user to set the initial state of the model. The user must provide a vector of proportions indicating the following values: (1) proportion of non-infected agents already removed, and (2) proportion of non-ifected agents already deceased.
See also
epiworld-methods
Other Models:
ModelDiffNet(),
ModelSEIR(),
ModelSEIRCONN(),
ModelSEIRD(),
ModelSEIRDCONN(),
ModelSEIRMixing(),
ModelSEIRMixingQuarantine(),
ModelSIR(),
ModelSIRCONN(),
ModelSIRD(),
ModelSIRLogit(),
ModelSIRMixing(),
ModelSIS(),
ModelSISD(),
ModelSURV()
Examples
model_sirdconn <- ModelSIRDCONN(
name = "COVID-19",
n = 100000,
prevalence = 0.01,
contact_rate = 5,
transmission_rate = 0.4,
recovery_rate = 0.5,
death_rate = 0.1
)
# Running and printing
run(model_sirdconn, ndays = 100, seed = 1912)
#> _________________________________________________________________________
#> Running the model...
#> ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||| done.
model_sirdconn
#> ________________________________________________________________________________
#> Susceptible-Infected-Removed-Deceased (SIRD) (connected)
#> It features 100000 agents, 1 virus(es), and 0 tool(s).
#> The model has 4 states.
#> The final distribution is: 3537 Susceptible, 0 Infected, 87030 Recovered, and 9433 Deceased.
plot(model_sirdconn, main = "SIRDCONN Model")